DNA Synthesis
The discovery of the double-helical nature of DNA by Watson & Crick explained how genetic information could be duplicated and passed on to succeeding generations. The strands of the double helix can separate and serve as templates for the synthesis of daughter strands. In conservative replication the two daughter strands would go to one daughter cell and the two parental strands would go to the other daughter cell. In semiconservative replication one parental and one daughter strand would go to each of the daughter cells.
Through experimentation it was determined that DNA replicates via a semiconservative mechanism. There are three possible mechanisms that can explain DNA's semiconservative replication.
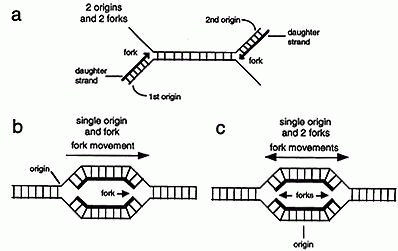
(a) DNA synthesis starts at a specific place on a chromosome called an origin. In the first mechanism one daughter strand is initiated at an origin on one parental strand and the second is initiated at another origin on the opposite parental strand. Thus only one strand grows from each origin. Some viruses use this type of mechanism.
(b) In the second mechanism replication of both strands is initiated at one origin. The site at which the two strands are replicated is called the replication fork. Since the fork moves in one direction from the origin this type of replication is called unidirectional. Some types of bacteria use this type of mechanism.
(c) In the third mechanism two replication forks are initiated at the origin and as synthesis proceeds the two forks migrate away from one another. This type of replication is called bi-directional. Most organisms, including mammals, use bi-directional replication.

Requirements for DNA Synthesis
There are four basic components required to initiate and propagate DNA synthesis. They are: substrates, template, primer and enzymes.
Substrates
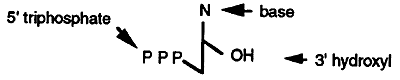
Four deoxyribonucleotide triphosphates (dNTP's) are required for DNA synthesis (note the only difference between deoxyribonucleotides and ribonucleotides is the absence of an OH group at position 2' on the ribose ring). These are dATP, dGTP, dTTP and dCTP. The high energy phosphate bond between the a and b phosphates is cleaved and the deoxynucleotide monophosphate is incorporated into the new DNA strand.
Ribonucleoside triphosphates (NTP's) are also required to initiate and sustain DNA synthesis. NTP's are used in the synthesis of RNA primers and ATP is used as an energy source for some of the enzymes needed to initiate and sustain DNA synthesis at the replication fork.
Template
The nucleotide that is to be incorporated into the growing DNA chain is selected by base pairing with the template strand of the DNA. The template is the DNA strand that is copied into a complementary strand of DNA.
Primer
The enzyme that synthesizes DNA, DNA polymerase, can only add nucleotides to an already existing strand or primer of DNA or RNA that is base paired with the template.
Enzymes
An enzyme, DNA polymerase, is required for the covalent joining of the incoming nucleotide to the primer. To actually initiate and sustain DNA replication requires many other proteins and enzymes which assemble into a large complex called a replisome. It is thought that the DNA is spooled through the replisome and replicated as it passes through.
DNA Synthesis, 5' to 3'
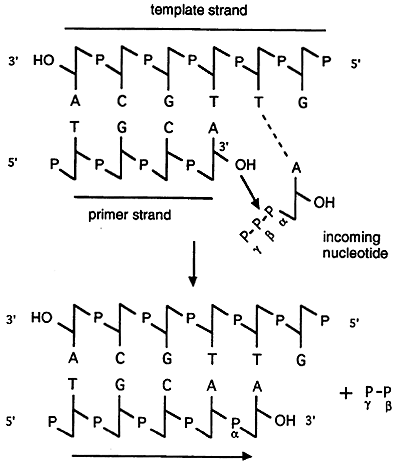
The major catalytic step of DNA synthesis is shown below. Notice that DNA synthesis always occurs in a 5' to 3' direction and that the incoming nucleotide first base pairs with the template and is then linked to the nucleotide on the primer.
DNA Synthesis is Semidiscontinuous
Since all known DNA polymerases can synthesize only in a 5' to 3' direction a problem arises in trying to replicate the two strands of DNA at the fork.
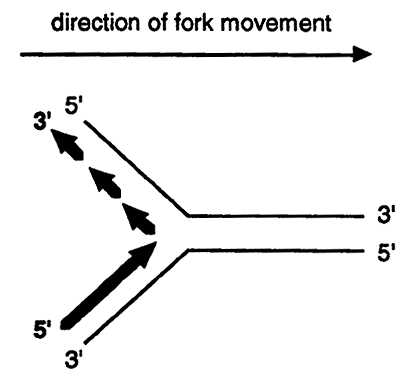
Notice that the top strand must be discontinuously replicated in short stretches thus the replication of both parental strands is a semidiscontinuous process. The strand that is continuously synthesized is called the leading strand while the strand that is discontinuously synthesized is called the lagging strand.
Leading Strand Synthesis
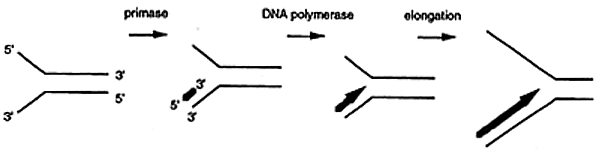
DNA synthesis requires a primer usually made of RNA. A primase synthesizes the ribonucleotide primer ranging from 4 to 12 nucleotides in length. DNA polymerase then incorporates a dNMP onto the 3' end of the primer initiating leading strand synthesis. Only one primer is required for the initiation and propagation of leading strand synthesis.
Lagging Strand Synthesis
Lagging strand synthesis is much more complex and involves five steps.
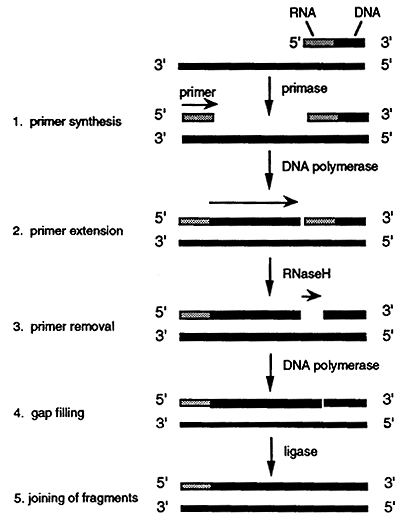
1. As the leading strand is synthesized along the lower parental strand the top parental strand becomes exposed. The strand is then recognized by a primase which synthesizes a short RNA primer.
2. DNA polymerase then incorporates a dNMP onto the 3" end of the primer and initiates lagging strand synthesis. The polymerase extends the primer for about 1,000 nucleotides until it comes in contact with the 5' end of the preceding primer. These short segments of RNA/DNA are known as Okazaki fragments.
3. When the DNA polymerase encounters the preceding primer it dissociates. The RNA is then removed by a specialized DNA polymerase or by an enzyme called RNaseH. Ribonucleotides are then excised one at a time in a 5' to 3' direction. The RNaseH leaves a phosphate group at the 5' end of the adjoining DNA segment thus leaving a gap.
4. The gap is filled by a DNA polymerase which uses an Okazaki fragment as a primer.
5. The 3' hydroxyl group on the 3' nucleotide terminus is then covalently joined, using DNA ligase, to the free 5' phosphate of the previously made lagging segment.
Structure of DNA
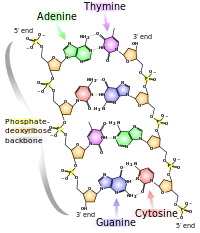
DNA Polymerases
There are many types of DNA polymerases which can excise, fill gaps, proofread, repair and replicate.
Other Factors Required for DNA Synthesis
Origins: Origins are unique DNA sequences that are recognized by a protein that builds the replisome. Origins have been found in bacterial, plasmid, viral, yeast and mitochondrial DNA and have recently been discovered in mammalian DNA. Specific origins are used for initiating DNA replication in humans. Most origins have a site that is recognized and bound by an origin-binding protein. When the origin-binding protein binds to the origin the A + T rich sequence becomes partially denatured allowing other replication factors known as cis-acting factors to bind and initiate DNA replication.
Origin-binding Protein: binds and partially denatures the origin DNA while binding to another enzyme called helicase.
Helicases: unwind double stranded DNA.
Single-stranded DNA Binding Protein (SSB): enhances the activity of the helicase and prevents the unwound DNA from renaturing.
Primase: synthesize the RNA primers required for initiating leading and lagging strand synthesis.
DNA Polymerase: recognizes the RNA primers and extends them in the 5' to 3' direction.
Processivity Factors: help load the polymerase onto the primer-template while anchoring the polymerase to the DNA.
Topoisomerase: removes the positive supercoils that form as the fork is unwound by the helicase.
RNaseH: removes RNA portions from Okazaki fragments.
Ligase: seals the nicks after filling in the gaps left by DNA polymerase.
Coordination of Leading and Lagging Strand Synthesis
Leading and lagging strand synthesis is thought to be coordinated at a replication fork. The two polymerases are held together by another set of proteins, tg, which are near the fork that is being unwound and simultaneously primed by helicase-primase. Both polymerases are bound by a processivity factor, b. Upon completing an Okazaki fragment the lagging strand polymerase release the b factor and dissociates from the DNA. The tg complex then loads the new b factor/primer complex onto the lagging strand polymerase which initiates a new round.....
Telomerase
Leading strand synthesis can proceed all the way to the end of a chromosome however lagging strand synthesis can not. Consequently the 3' tips of each daughter chromosome would not be replicated.
Telomerase ( also AKA telomere terminal transferase) extends the 3' ends of a chromosome by adding numerous repeats of a six base pair sequence until the 3' end of the lagging strand is long enough to be primed and extended by DNA polymerase.
Telomerase recognizes the tips of chromosomes also know as telomeres. The DNA sequences of telomeres have been determined in several organisms and consist of numerous repeats of a 6 to 8 base long sequence, [TTGGGG]n.
Telomeres have been found to progressively shorten in certain types of cells. These cells appear to lack Telomerase activity. When telomeric length shortens to a critical point the cell dies. Cells derived from rapidly proliferating tissues, such as tumors, have telomeres that are unusually long. This indicates that Telomerase activity may be necessary for the proliferation of tumor cells. Telomerase activity is found in ovarian cancer cells but not in normal ovarian tissue. Thus it may be possible to develop anti-tumor drugs that function to inhibit telomerase activity.
Chemical Inhibitors of DNA Replication
Some types of drugs function by inhibiting DNA replication.
Substrate Analogs: analogs of dNTP's which function as chain terminators can be incorporated into DNA. These analogs are usually either missing the 3' hydroxyl group or have a chemical group, other than hydroxyl, in the 3' position.
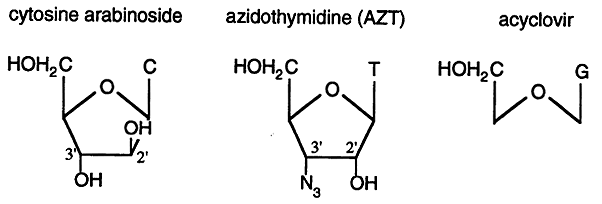
Cytosine Arabinoside: is an anticancer drug used to treat leukemia.
Azidothymidine (AZT): was used as an anti-HIV drug that, while effective in tissue culture experiments, proved to be ineffective for treating HIV in humans.
Acyclovir: is an effective anti-herpes virus drug.
Intercalating Agents: are compounds with fused aromatic ring systems that can wedge (intercalate) between the stacked base pairs of DNA. This disrupts the structure of the DNA so that the replicative enzymes have difficulty in synthesizing DNA past the "intercalated" sites. Anthracycline glycosides and Actinomycin D are intercalators used to treat a variety of cancers.
DNA Damaging Agents: a variety of compounds such as Cisplatin, cause chemical damage to DNA and are used in the treatment of cancers.
Topoisomerase Inhibitors: Nalidixic acid and Fluoroquinolones are antibiotics used to inhibit bacterial topoisomerases.
DNA Mutation and Repair
A mutation, which may arise during replication and/or recombination, is a permanent change in the nucleotide sequence of DNA. Damaged DNA can be mutated either by substitution, deletion or insertion of base pairs. Mutations, for the most part, are harmless except when they lead to cell death or tumor formation. Because of the lethal potential of DNA mutations cells have evolved mechanisms for repairing damaged DNA.
Types of Mutations
There are three types of DNA Mutations: base substitutions, deletions and insertions.
1. Base Substitutions
Single base substitutions are called point mutations, recall the point mutation Glu -----> Val which causes sickle-cell disease. Point mutations are the most common type of mutation and there are two types.
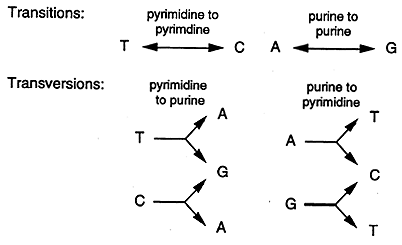
Transition: this occurs when a purine is substituted with another purine or when a pyrimidine is substituted with another pyrimidine.
Transversion: when a purine is substituted for a pyrimidine or a pyrimidine replaces a purine.
Point mutations that occur in DNA sequences encoding proteins are either silent, missense or nonsense.
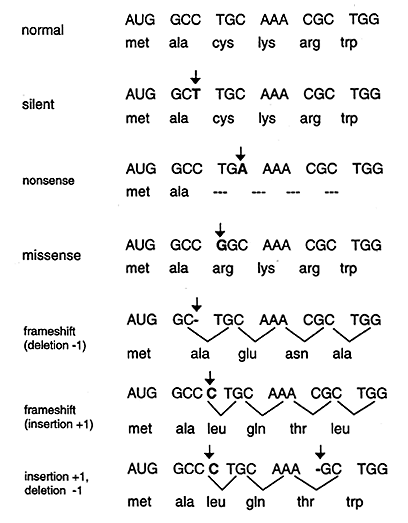
Silent: If abase substitution occurs in the third position of the codon there is a good chance that a synonymous codon will be generated. Thus the amino acid sequence encoded by the gene is not changed and the mutation is said to be silent.
Missence: When base substitution results in the generation of a codon that specifies a different amino acid and hence leads to a different polypeptide sequence. Depending on the type of amino acid substitution the missense mutation is either conservative or nonconservative. For example if the structure and properties of the substituted amino acid are very similar to the original amino acid the mutation is said to be conservative and will most likely have little effect on the resultant proteins structure / function. If the substitution leads to an amino acid with very different structure and properties the mutation is nonconservative and will probably be deleterious (bad) for the resultant proteins structure / function (i.e. the sickle cell point mutation).
Nonsense: When a base substitution results in a stop codon ultimately truncating translation and most likely leading to a nonfunctional protein.
2. Deletions
A deletion, resulting in a frameshift, results when one or more base pairs are lost from the DNA (see Figure above). If one or two bases are deleted the translational frame is altered resulting in a garbled message and nonfunctional product. A deletion of three or more bases leave the reading frame intact. A deletion of one or more codons results in a protein missing one or more amino acids. This may be deleterious or not.
3. Insertions
The insertion of additional base pairs may lead to frameshifts depending on whether or not multiples of three base pairs are inserted. Combinations of insertions and deletions leading to a variety of outcomes are also possible.
Causes of Mutations
Errors in DNA Replication
On very, very rare occasions DNA polymerase will incorporate a noncomplementary base into the daughter strand. During the next round of replication the missincorporated base would lead to a mutation. This, however, is very rare as the exonuclease functions as a proofreading mechanism recognizing mismatched base pairs and excising them.
Errors in DNA Recombination
DNA often rearranges itself by a process called recombination which proceeds via a variety of mechanisms. Occasionally DNA is lost during replication leading to a mutation.
Chemical Damage to DNA
Many chemical mutagens, some exogenous, some man-made, some environmental, are capable of damaging DNA. Many chemotherapeutic drugs and intercalating agent drugs function by damaging DNA.
Radiation
Gamma rays, X-rays, even UV light can interact with compounds in the cell generating free radicals which cause chemical damage to DNA.
DNA Repair
Damaged DNA can be repaired by several different mechanisms.
Mismatch Repair
Sometimes DNA polymerase incorporates an incorrect nucleotide during strand synthesis and the 3' to 5' editing system, exonuclease, fails to correct it. These mismatches as well as single base insertions and deletions are repaired by the mismatch repair mechanism. Mismatch repair relies on a secondary signal within the DNA to distinguish between the parental strand and daughter strand, which contains the replication error. Human cells posses a mismatch repair system similar to that of E. coli, which is described here. Methylation of the sequence GATC occurs on both strands sometime after DNA replication. Because DNA replication is semi-conservative, the new daughter strand remains unmethylated for a very short period of time following replication. This difference allows the mismatch repair system to determine which strand contains the error. A protein, MutS recognizes and binds the mismatched base pair.
Another protein, MutL then binds to MutS and the partially methylated GATC sequence is recognized and bound by the endonuclease, MutH. The MutL/MutS complex then links with MutH which cuts the unmethylated DNA strand at the GATC site. A DNA Helicase, MutU unwinds the DNA strand in the direction of the mismatch and an exonuclease degrades the strand. DNA polymerase then fills in the gap and ligase seals the nick. Defects in the mismatch repair genes found in humans appear to be associated with the development of hereditary colorectal cancer.
Nucleotide Excision Repair (NER)
NER in human cells begins with the formation of a complex of proteins XPA, XPF, ERCC1, HSSB at the lesion on the DNA. The transcription factor TFIIH, which contains several proteins, then binds to the complex in an ATP dependent reaction and makes an incision. The resulting 29 nucleotide segment of damaged DNA is then unwound, the gap is filled (DNA polymerase) and the nick sealed (ligase).
Direct Repair of Damaged DNA
Sometimes damage to a base can be directly repaired by specialized enzymes without having to excise the nucleotide.
Recombination Repair
This mechanism enables a cell to replicate past the damage and fix it later.
Regulation of Damage Control
DNA repair is regulated in mammalian cells by a sensing mechanism that detects DNA damage and activates a protein called p53. p53 is a transcriptional regulatory factor that controls the expression of some gene products that affect cell cycling, DNA replication and DNA repair. Some of the functions of p53, which are just being determined, are: stimulation of the expression of genes encoding p21 and Gaad45. Loss of p53 function can be deleterious, about 50% of all human cancers have a mutated p53 gene.
The p21 protein binds and inactivates a cell division kinase (CDK) which results in cell cycle arrest. p21 also binds and inactivates PCNA resulting in the inactivation of replication forks. The PCNA/Gaad45 complex participates in excision repair of damaged DNA.
Some examples of the diseases resulting from defects in DNA repair mechanisms.
Hereditary nonpolyposis colorectal cancer
© Dr. Noel Sturm 2021