Regulation of Gene Expression
Cellular function is influenced by cellular environment. Adaptation to specific environments is achieved by regulating the expression of genes that encode the enzymes and proteins needed for survival in a particular environment. Factors that influence gene expression include nutrients, temperature, light, toxins, metals, chemicals, and signals from other cells. Malfunctions in the regulation of gene expression can cause various human disorders and diseases.
Regulation in Prokaryotes
Bacteria have a simple general mechanism for coordinating the regulation of genes that encode products involved in a set of related processes. The gene cluster and promoter, plus additional sequences that function together in regulation are called an operon.
The Lactose Operon (lac operon)
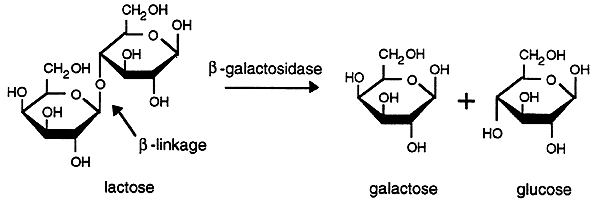
The lactose operon of E. coli encodes the enzyme b-galactosidase which hydrolyzes lactose into galactose and glucose.

The lac operon contains three cistrons or DNA fragments that encode a functional protein. The proteins encoded by cistrons may function alone or as sub-units of larger enzymes or structural proteins.

The Z gene encodes for b-galactosidase. The Y gene encodes a permease that facilitates the transport of lactose into the bacterium. The A gene encodes a thiogalactoside transacetylase whose function is not known. All three of these genes are transcribed as a single, polycistronic mRNA. Polycistronic RNA contains multiple genetic messages each with its own translational initiation and termination signals.
Regulation of the lac Operon
The activity of the promoter that controls the expression of the lac operon is regulated by two different proteins. One of the proteins prevents the RNA polymerase from transcribing (negative control), the other enhances the binding of RNA polymerase to the promoter (positive control).
Negative Control of the lac Operon
The protein that inhibits transcription of the lac operon is a tetramer with four identical subunits called lac repressor. The lac repressor is encoded by the lacI gene, located upstream of the lac operon and has its own promoter. Expression of the lacI gene is not regulated and very low levels of the lac repressor are continuously synthesized. Genes whose expression is not regulated are called constitutive genes.
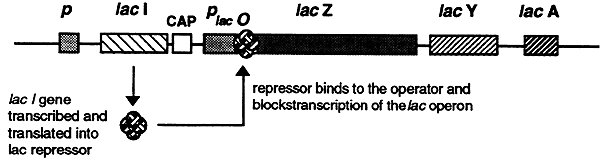
In the absence of lactose the lac repressor blocks the expression of the lac operon by binding to the DNA at a site, called the operator that is downstream of the promoter and upstream of the transcriptional initiation site. The operator consists of a specific nucleotide sequence that is recognized by the repressor which binds very tightly, physically blocking (strangling) the initiation of transcription.
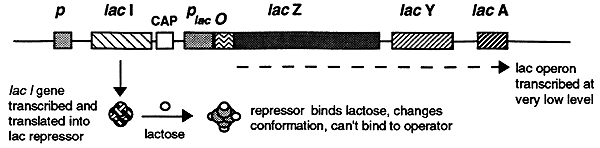
The lac repressor has a high affinity for lactose. When a small amount of lactose is present the lac repressor will bind it causing dissociation from the DNA operator thus freeing the operon for gene expression. Substrates that cause repressors to dissociate from their operators are called inducers and the genes that are regulated by such repressors are called inducible genes.
Positive Control of the lac Operon
Although lactose can induce the expression of lac operon, the level of expression is very low. The reason for this is that the lac operon is subject to catabolite repression or the reduced expression of genes brought on by growth in the presence of glucose. Glucose is very easily metabolized so is the preferred fuel source over lactose, hence it makes sense to prevent expression of lac operon when glucose is present.
The strength of a promoter is determined by its ability to bind RNA polymerase and to form an open complex. The promoter for the lac operon is weak and consequently the lac operon is poorly transcribed upon induction. There is a binding site, upstream from the promoter, for a protein called the catabolite activator protein (CAP). When the CAP protein binds it distorts the DNA so that the RNA polymerase can bind more effectively, thus transcription of the lac operon is greatly enhanced. In order to bind the CAP must first bind cyclic AMP (cAMP), a second messenger synthesized from ATP by the enzyme Adenylate Cyclase.
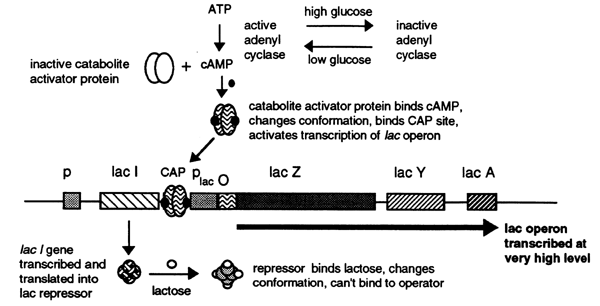
In the presence of glucose circulating cAMP levels are very low and consequently the initiation of transcription from the lac operon is very low. As glucose levels decrease the concentration of cAMP increases activating CAP which in turn binds to the CAP site stimulating transcription. The cAMP-CAP complex is called a positive regulator.
The Arabinose Operon
Arabinose is a five-carbon sugar that can serve as an energy and carbon source for E. coli. Arabinose must first be converted into ribulose-5-phosphate before it can be metabolized. The arabinose operon has three genes,araB, araA and araD that encode for three enzymes to carry out this conversion. A fourth gene, araC, which has its own promoter, encodes a regulatory factor called the C protein.
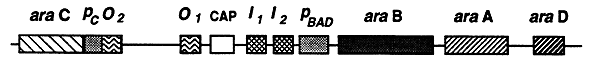
The regulatory sites of the ara operon include four sites that bind the C protein and one CAP binding site. The araO1 and araO2 sites are upstream of the promoter and CAP binding sites. The other two C protein binding sites called araI1 and araI2 are located between the CAP binding site and the promoter.
Negative Control of the araC Operon
In the absence of arabinose, dimers of the C protein bind to araO2, araO1 and araI1. The C proteins bound to araO2 and araI1 associate with one another causing the DNA between them to form a loop effectively blocking transcription of the operon.
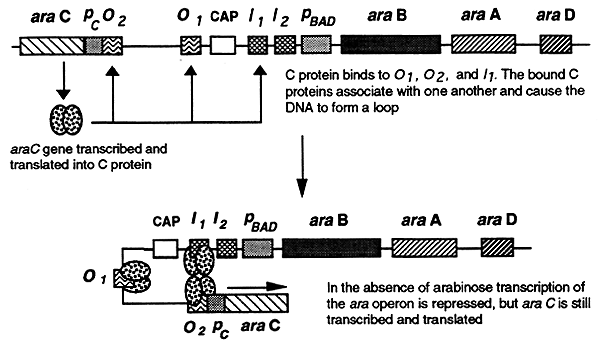
Positive Control of the araC Operon
The C protein binds arabinose and undergoes a conformational change that enables it to also bind the araO2 and araI2 sites. This results in the generation of a different DNA loop that is formed by the interaction of C proteins bound to the araO1 and araO2 sites.
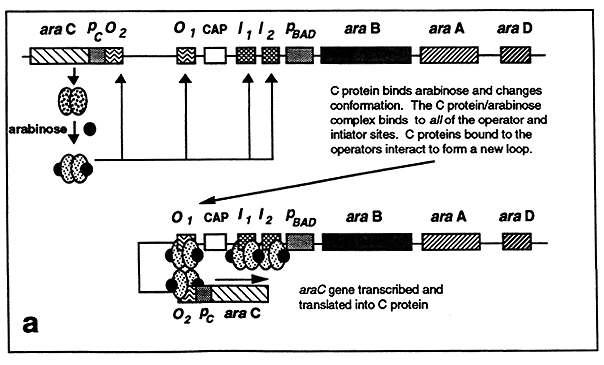
The formation of this loop stimulates transcription of the araC gene resulting in additional C protein synthesis, thus the C protein autoregulates its own synthesis. In the absence of glucose, cAMP-CAP is formed which binds to the CAP site. C protein bound at the araI1 and araI2 sites interacts with the bound CAP enabling RNA polymerase to initiate transcription from the ara operon promoter.
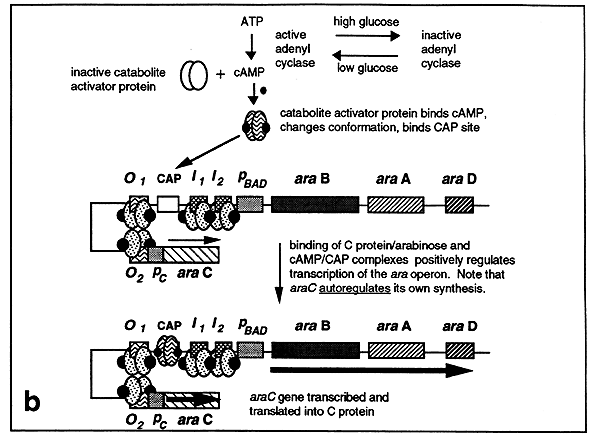
The Tryptophan Operon
E. coli can synthesize all 20 of the natural amino acids. Amino acid synthesis consumes a lot of energy, so to avoid wasting energy the operons that encode for amino acid synthesis are tightly regulated. The trp operon consists of five genes, trpE, trpD, trpC, trpB and trpA, that encode for the enzymes required for the synthesis of tryptophan.
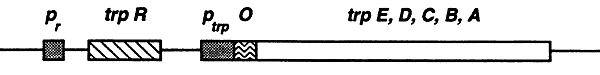
The trp operon is regulated by two mechanisms, negative corepression and attenuation. Most of the operons involved in amino acid synthesis are regulated by these two mechanisms.
Negative Corepression
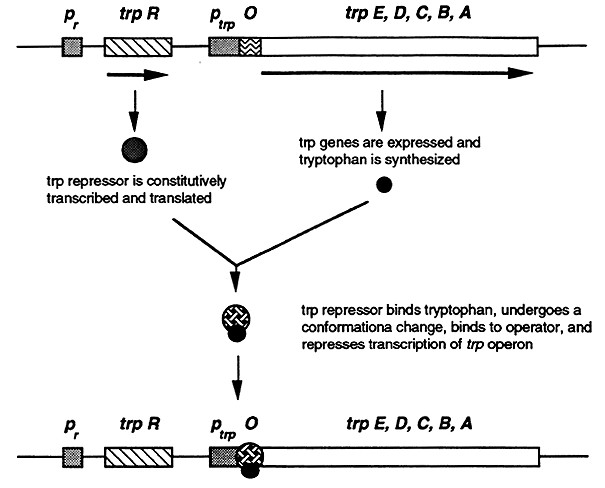
The trp operon is negatively controlled by the trp repressor, a product of the trpR gene. The trp repressor binds to the operator and blocks transcription of the operon. However, in order to bind to the operator the repressor must first bind to Trp hence tryptophan is a corepressor. In the absence of Trp the trp repressor dissociates and transcription of the trp operon is initiated.
Attenuation
Attenuation regulates the termination of transcription as a function of tryptophan concentration. At low levels of trp full length mRNA is made, at high levels transcription of the trp operon is prematurely halted. Attenuation works by coupling transcription to translation. Prokaryotic mRNA does not require processing and since prokaryotes have no nucleus translation of mRNA can start before transcription is complete. Consequently regulation of gene expression via attenuation is unique to prokaryotes.
a. Attenuation is mediated by the formation of one of two possible stem-loop structures in a 5' segment of the trp operon in the mRNA.
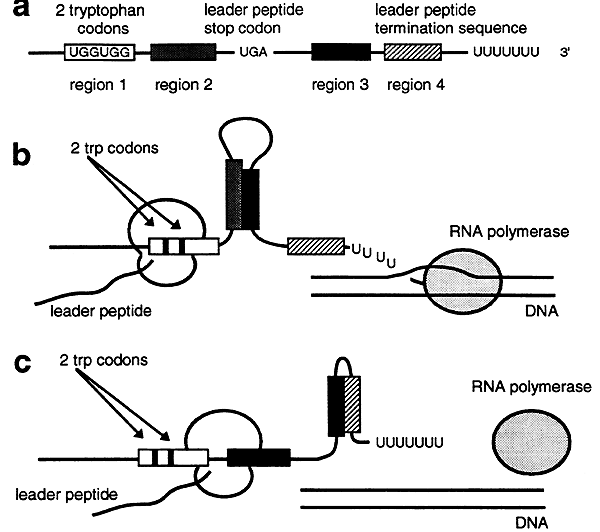
b. If tryptophan concentrations are low then translation of the leader peptide is slow and transcription of the trp operon outpaces translation. This results in the formation of a nonterminating stem-loop structure between regions 2 and 3 in the 5' segment of the mRNA. Transcription of the trp operon is then completed.
c. If tryptophan concentrations are high the ribosome quickly translates the mRNA leader peptide. Because translation is occurring rapidly the ribosome covers region 2 so that it can not attach to region 3. Consequently the formation of a stem-loop structure between regions 3 and 4 occurs and transcription is terminated.
Regulation of Gene Expression in Eukaryotes
The genetic information of a human cell is a thousand fold greater than that of a prokaryotic cell. Things are further complicated by the number of cell types and the fact that each cell type must express a particular subset of genes at different points in an organisms development. Regulating gene expression so that a particular subset of genes is expressed in a specific tissue at specific points of development is very complicated. This increased complexity in regulation lends itself to malfunctions that cause disease. Three ways that eukaryotes regulate gene expression will be discussed: alteration of gene content or position, transcriptional regulation and alternative RNA processing.
1. Alteration of Gene Content or Position
The copy number of a gene or its location on the chromosome can greatly effect its level of expression. Gene content or location can be altered by gene amplification, diminution or rearrangement.
Gene Amplification
The expression of a particular gene can be augmented by amplifying its copy number. Histone proteins and rRNA are needed in large quantities by almost all eukaryotic cells therefore the genes encoding histones and rRNA exist in a permanently amplified state. Gene amplification can present problems with the use of chemotherapeutic drugs. Methotrexate inhibits dihydrofolate reductase, the enzyme responsible for regenerating the folates used in nucleotide synthesis. Tumor cells often become resistant to the drug because the gene encoding dihydrofolate reductase is amplified by several hundred fold resulting in more enzyme production then the drug can handle.
Gene Diminution
A gene whose expression is only needed at a particular developmental point or in a particular tissue may be shut off by gene diminution. As reticulocytes mature into red blood cells all of their genes are lost as the nucleus is degraded.
Gene Rearrangements
Gene rearrangement is used to generate each of the genes encoding the millions of different antibodies that are produced by B cells. Sometimes bad gene rearrangements occur that lead to improper gene regulation. This frequently occurs in cancer cells. Translocation of a segment from chromosome 8 to chromosomes that encode immunoglobulins leads to activation of a gene that transforms healthy B cells into Burkitt's lymphoma cells (unregulated proliferating B cells).
2. Transcriptional Regulation
Through Chromosomal Packaging
Regions of each of the different chromosomes are either packaged as heterochromatin or euchromatin. In heterochromatin the DNA is very tightly condensed and rendered inaccessible to the transcriptional machinery, consequently heterochromatin is transcriptionally inactive. In human females one of each of the two X chromosomes is completely inactivated by being packaged into a heterochromatin to form a Barr body. The Cys residues in DNA in the heterochromatin are heavily methylated suggesting that methylation may play a role in the maintenance of heterochromatin. Drugs that interfere with methylation cause activation of previously inactive genes found in heterochromatin.
In euchromatin the DNA is not as condensed and is accessible to the transcription machinery. The regions of a chromosome that are maintained as hetero- and eu- chromatin may vary in a cell specific manner. This may enable the cells of specific tissues to express a particular subset of genes required for tissue function.
Through Individual Genes
Trans-acting Elements
Proteins that participate in regulating gene expression are often called trans acting elements. At least 100 different proteins, many specific for the regulation of a particular gene, are known. Others play a more general role in regulating gene expression in a manner analogous to the activation of numerous prokaryotic genes by the CAP-cAMP complex. Trans-acting factors have multiple domains required for activity and may include DNA-binding, transcription-activating and ligand-binding domains.
DNA Binding Domains
DNA binding domains recognize specific DNA sequences in the regulatory regions of a gene. The DNA-binding domains of a regulatory protein generally consist of one of three motifs: helix-turn-helix, zinc finger or leucine zipper. DNA-binding proteins possessing these motifs bind with high affinity to their recognition sites and with low affinity to other DNA. A very small portion of the protein makes contact with the DNA through H-bonds and van der Waals interactions between amino acid side chains and the functional groups in the major groove and the phosphate backbone of the DNA. The remainder of the protein is involved in proper positioning of the DNA-binding domain and in making protein-protein contacts with other transcriptional proteins.
The Helix-Turn-Helix Motif
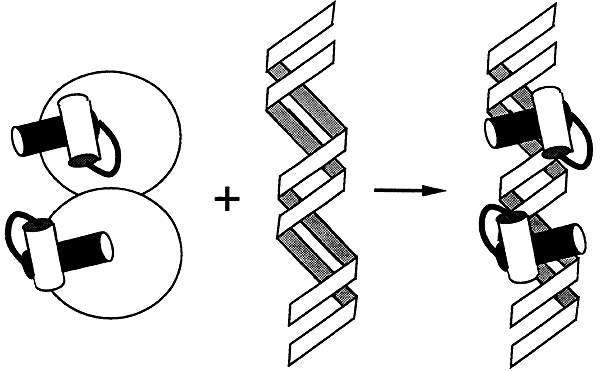
Proteins with this motif form symmetric dimers that recognize a symmetric palindromic DNA sequence. Each monomer of the dimer contains a region in which two a helices are held at 90 degrees to each other by a turn of four amino acids. One set of helices makes contact with about five base pairs in the major groove. The other set sits atop the phosphate backbone and helps to properly position the set of helices that fits into the major groove.
The Zinc-Finger Motif
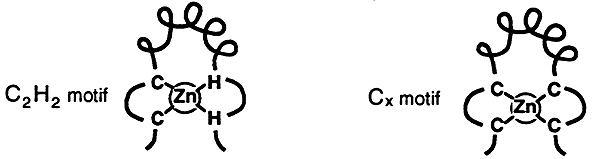
Proteins possessing this motif contain between 2 to 9 repeated domains that are each centered on a tetrahedrally coordinated zinc ion. Each zinc coordinated domain forms a loop containing an a-helix, this loop is called a zinc-finger. There are two types of zinc fingers: the C2H2 finger and the Cx finger.
C2H2 Finger:
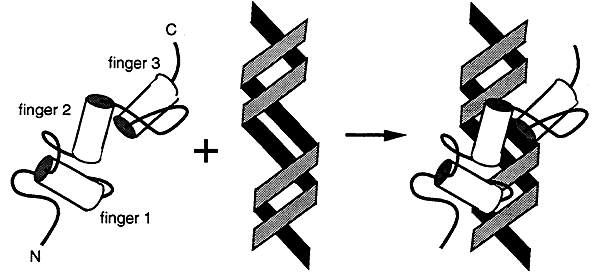
Three fingers interact with the major groove and wrap around the DNA. Many transcription factors have this type of domain.
Cx Finger:
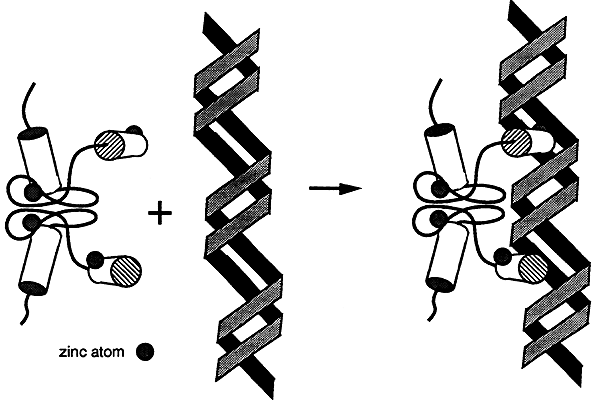
Proteins with this motif bind as dimers to the major groove of the DNA. Many steroid receptors have this type of domain.
The Leucine Zipper Motif
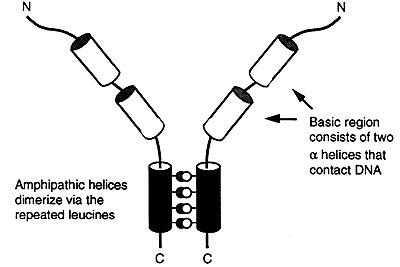
Proteins with this type of motif have an amphipathic a-helix at their carboxyl terminus. One side of the helix consists of hydrophobic groups, usually leucine, that are repeated every seventh position for several turns of the helix. The other face consists of charged and polar groups.
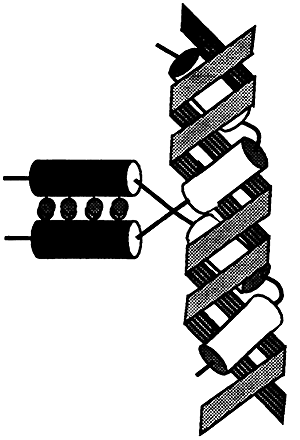
Proteins with this motif bind as dimers to the major groove of the DNA. The two a-helices of each arm enter the major groove and wrap around the double helix. Several oncogenes use this type of motif.
Transcription-Activating Domains
These domains generally act separately and independently of the DNA-binding domains. Transcription-activating domains enhance transcription by physically ineracting with other regulatory proteins and/or with RNA polymerase. The actual mechanisms by which these domains activate or enhance transcription are not known.
Ligand-Binding Domains
Steroid hormones, thyroid hormones and retinoic acid are examples of ligands that activate transcription by binding to a specific domain on a receptor protein. Upon binding the receptor undergoes a conformational change that enables it to bind DNA. Once bound to the DNA a receptor protein can activate or repress transcription of the target gene.
Cis-acting Elements
Cis-acting elements are DNA sequences that are recognized and bound by the trans-acting elements that regulate transcription. There are two major types of cis-acting elements: promoters and regulatory elements.
Promoters
Promoters are the sites where RNA polymerase must bind to the DNA in order to initiate transcription (see "RNA Synthesis and Processing" lecture). The rate or efficiency of promoter use by RNA polymerase is affected by the regulatory elements.
Regulatory Elements
Regulatory elements are specific DNA sequences that are recognized and bound by the trans-acting elements that stimulate or inhibit the expression of a particular gene. There are two types: enhancers and response elements.
Enhancers are regulatory elements that increase or repress the rate of gene transcription.
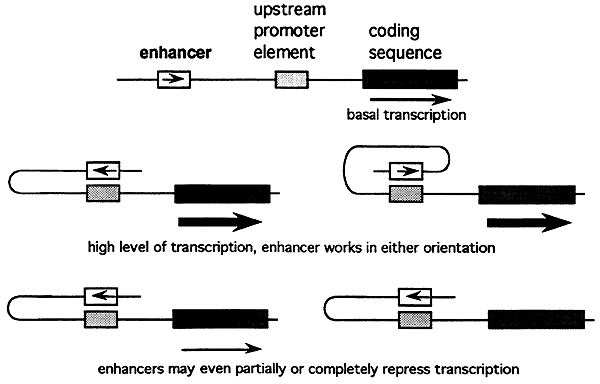
Response Elements are regulatory sequences that facilitate the coordinated regulation of a group of genes. Certain ligands such as steroid hormones and cAMP bind to their receptors which in turn bind to their response element to activate or inhibit transcription.
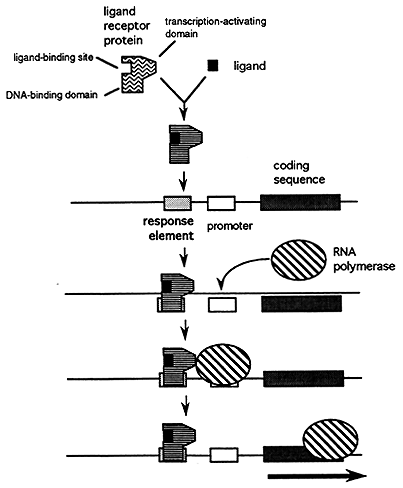
3. Alternative Processing
Alternative Start Sites
Initiating transcription at an alternative start site places a different exon at the 5' end of the transcript. Examples of genes that use alternative start sites as a form of regulation include amylase, myosin and alcohol dehydrogenase.
Alternative Polyadenylation Sites
Immunoglobin (antibody) heavy chains use an alternative polyadenylation site to affect the length of transcripts. The longer transcript encodes the mm form which is localized to the cell membranes of lymphocytes, the shorter transcript encodes the secreted form, ms.
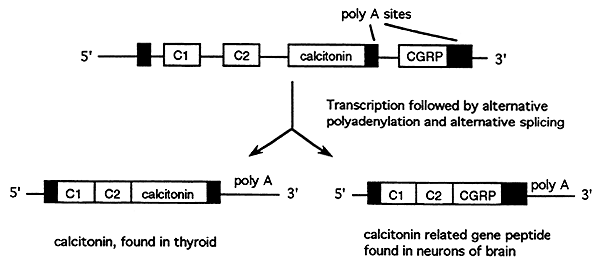
Alternative Splice Sites
Alternative splice sites are used to generate similar proteins with tissue specific functions called isoforms. Many peptide hormones exist as isoforms such as the calcitonin gene which is differentially spliced to produce calcitonin in the thyroid and calcitonin gene-related peptide in the neurons.
Regulation of mRNA Stability
The stability of mRNA is quite variable form gene to gene. These variations in stability govern the length of time that mRNA is available for translation and hence the amount of protein that is synthesized. The half-lives of mRNA vary from 10 hours to minutes. Sequences in the 3' untranslated region of mRNA which serve as signals for rapid degradation have been identified in some mRNA's with very short half-lives. The length of the poly A tail also affects mRNA stability, with longer tails tending to have longer half-lives.
© Dr. Noel Sturm 2020